Personal genomics
This article needs more reliable medical references for verification or relies too heavily on primary sources. (April 2012) |  |
| Part of a series on |
| Genetic genealogy |
|---|
| Concepts |
| Related topics |
Personal genomics is the branch of genomics concerned with the sequencing and analysis of the genome of an individual. The genotyping stage employs different techniques, including single-nucleotide polymorphism (SNP) analysis chips (typically 0.02% of the genome), or partial or full genome sequencing. Once the genotypes are known, the individual's genotype can be compared with the published literature to determine likelihood of trait expression and disease risk.
Automated sequencers have increased the speed and reduced the cost of sequencing, making it possible to offer genetic testing to consumers.
Use of personal genomics in predictive and precision medicine
Predictive medicine is the use of the information produced by personal genomics techniques when deciding what medical treatments are appropriate for a particular individual. Precision medicine is focused on "a new taxonomy of human disease based on molecular biology" [1]
Examples of the use of predictive and precision medicine include inherited medical genomics, cancer genomics and pharmacogenomics. In pharmacogenomics genetic information can be used to select the most appropriate drug to prescribe to a patient. The drug should be chosen to maximize the probability of obtaining the desired result in the patient and minimize the probability that the patient will experience side effects. Genetic information may allow physicians to tailor therapy to a given patient, in order to increase drug efficacy and minimize side effects. As of Oct 2012 there are 167 examples of drug gene pairs for which this information is currently useful in clinical practice and this number has been growing rapidly.[2]
Disease risk may be calculated based on genetic markers and genome-wide association studies for common medical conditions, which are multifactorial and include environmental components in the assessment. Diseases which are individually rare (less than 200,000 people affected in the USA) are nevertheless collectively common (affecting roughly 8-10% of the US population[3]). Over 2500 of these diseases (including a few more common ones) have predictive genetics of sufficiently high clinical impact that they are recommended as medical genetic tests available for single genes (and in whole genome sequencing) and growing at about 200 new genetic diseases per year.[4]
Cost of sequencing an individual's genome
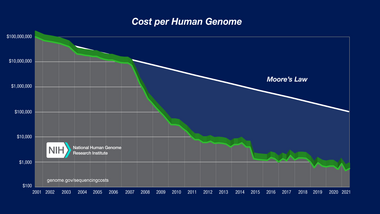
The cost of sequencing a human genome is dropping rapidly, due to the continual development of new, faster, cheaper DNA sequencing technologies such as "next generation DNA sequencing".
The National Human Genome Research Institute, part of the U.S. National Institutes of Health, has set a target to be able to sequence a human-sized genome for US$100,000 by 2009 and US$1,000 by 2014.[5][6]
There are 6 billion base pairs in the diploid human genome. Statistical analysis reveals that a coverage of approximately ten times is required to get coverage of both alleles in 90% human genome from 25 base pair reads with shotgun sequencing.[7] This means a total of 60 billion base pairs that must be sequenced. An Applied Biosystems SOLiD, Illumina or Helicos[8] sequencing machine can sequence 2 to 10 billion base pairs in each $8,000 to $18,000 run. The cost must also take into account personnel costs, data processing costs, legal, communications and other costs. One way to assess this is via commercial offerings. The first such whole diploid genome sequencing (6 billion bp, 3 billion from each parent) was from Knome and their price dropped from $350,000 in 2008 to $99,000 in 2009.[9][10] This inspects 3000-fold more bases of the genome than SNP chip-based genotyping, identifying both novel and known sequence variants, some relevant to personal health or ancestry.[11] In June 2009, Illumina announced the launch of its own Personal Full Genome Sequencing Service at a depth of 30X for $48,000 per genome.[12] In 2010, they cut the price to $19,500.[13]
In 2009, Complete Genomics of Mountain View announced that it would provide full genome sequencing for $5,000, from June 2009.[14] This will only be available to institutions, not individuals.[15] Prices are expected to drop further over the next few years through economies of scale and increased competition.[16][17] As of 2014, full exome sequencing was offered by Gentle for less than $2,000, including personal counseling along with the results. [18]
Projects and services already available
- Sequencing.com provides free, unlimited data storage for all genetic data[19] in high-security data centers[20] and software applications to analyze the data based on their patent-pending Real-Time Personalization™ technology.[21] The company invented an open API that translates genetic code into software code so that third-party developers without any training in genetics can integrate genetic code into their own apps.[22] The company also runs the not-for-profit Altruist Endeavor, which is an open data initiative consisting of a free, publically accessible online repository of anonymous genetic data in order to enable genetic research.[23]
- The Genographic Project is a project of the National Geographic Society and IBM to collect DNA samples to map historical human migration patterns. Launched in 2005, with 500,000 public participants as of December 2012, it helped to create the direct-to-consumer (DTC) genetic testing industry.
- The Personal Genome Project (PGP) is a long term, large cohort study based at Harvard Medical School which aims to sequence and publicize the complete genomes and medical records of 100,000 volunteers, in order to enable research into personal genomics and personalized medicine.
- SNPedia is a wiki that collects and shares information about the consequences of DNA variations, and through the associated program Promethease, anyone who has obtained DNA data about themselves (from any company) can get a free, independent report containing risk assessments and related information.
- deCODEme.com charged $1100 to carry out genotyping of approximately 1 million SNPs and provided risk estimates for 47 diseases as well as ancestry analyses. However, sales of genetic scans direct to consumer through deCODEme have now been discontinued.
- Navigenics began offering SNP-based genomic risk assessments as of April 2008. Navigenics is medically focused and emphasizes a clinician's and genetic counselor's role in interpreting results. Affymetrix Genome-Wide Human SNP Array 6.0, which genotypes 900,000 SNPs.
- Pathway Genomics analyzes over 100 genetic markers to identify genetic risk for common health conditions such as melanoma, prostate cancer and rheumatoid arthritis.
- 23andMe sells mail order kits for SNP genotyping The information is stored in a user profile and used to estimate the genetic risk of the consumer for over 240 diseases and conditions, as well as ancestry analysis. 23andMe utilizes a DNA array manufactured by Illumina.
- Illumina, Oxford Nanopore Technologies, Sequenom, Pacific Biosciences, Complete Genomics, and 454 Life Sciences are commercializing full genome sequencing but do not provide genetic analysis or counselling.[24][25][26][27][28]
- Gene by Gene provides Whole Genome Sequencing from $6995 to $7595, and other variants.
- Life Technologies
- Mapmygenome offers several SNP and NGS panels for personal genomics. See [2]
- Xcode Life Sciences offers actionable solutions for lifestyle conditions such as Diabetes, Obesity and Cardiovascular diseases. Nutrigenomics based recommendations are given to customers. Xcode uses a DNA array manufactured by Illumina. Xcode also has a skin genomics product that focuses on reducing/preventing wrinkles, acne and pigmentation. [3]
Ethical issues
Genetic discrimination is discriminating on the basis of information obtained from an individual’s genome. Genetic non-discrimination laws have been enacted in some US states[29] and at the federal level, by the Genetic Information Nondiscrimination Act (GINA). The GINA legislation prevents discrimination by health insurers and employers, but does not apply to life insurance or long-term care insurance. Given the ethical concerns about presymptomatic genetic testing of minors,[30][31][32][33] it is likely that personal genomics will first be applied to adults who can provide consent to undergo such testing, although genome sequencing is already proving valuable for children if any symptoms are present.[34]
Patients will need to be educated on interpreting their results and what they should be rationally taking from the experience. It is not only the average person who needs to be educated in the dimensions of their own genomic sequence but also professionals, including physicians and science journalists, who must be provided with the knowledge required to inform and educate their patients and the public [35] Examples of such efforts include the Personal Genetics Education Project (pgEd) and the Smithsonian collaboration with NHGRI[36]
Other issues
Full sequencing of the genome can identify polymorphisms that are so rare that no conclusions may be drawn about their impact, creating uncertainty in the analysis of individual genomes, particularly in the context of clinical care. Czech medical geneticist Eva Machácková writes: "In some cases it is difficult to distinguish if the detected sequence variant is a causal mutation or a neutral (polymorphic) variation without any effect on phenotype. The interpretation of rare sequence variants of unknown significance detected in disease-causing genes becomes an increasingly important problem."[37]
There is a heavy debate as to how relevant the results of personal genome kits are and whether or not the ramifications of knowing one’s predisposition to a disease is worth the potential psychological stress. There are also three potential problems associated with the validity of personal genome kits. The first issue is the test’s validity. Handling errors of the sample increases the likelihood for errors which could affect the test results and interpretation. The second affects the clinical validity, which could affect the test’s ability to detect or predict associated disorders. The third problem is the clinical utility of personal genome kits and associated risks, and the benefits of introducing them into clinical practices.[38]
Doctors are currently conducting tests for which some are not correctly trained to interpret the results. Many are unaware of how SNPs respond to one another. This results in presenting the client with potentially misleading and worrisome results which could strain the already overloaded health care system.[39] This may antagonize the individual to make uneducated decisions such as unhealthy lifestyle choices and family planning modifications. Moreover, negative results which may potentially be inaccurate, theoretically decrease the quality of life and mental health of the individual (such as increased depression and extensive anxiety).
There is also controversy regarding the concerns with companies testing individual DNA. There are issues such as "leaking" information, the right to privacy and what responsibility the company has to ensure this does not happen. Regulation rules are not clearly laid out. What is still not determined is who legally owns the genome information: the company or the individual whose genome has been read. There have been published examples of personal genome information being exploited.[40] Additional privacy concerns, related to, e.g., genetic discrimination, loss of anonymity, and psychological impacts, have been increasingly pointed out by the academic community [41] as well as government agencies.[42]
Conversely, sequencing one’s genome would allow for more personalized medical treatments using pharmacogenomics; the use of genetic information to select appropriate drugs.[43] Treatments can be catered to the individual and the certain genetic predispositions they may have (such as personalized chemotherapy).
Popular culture
The 1997 science fiction film GATTACA presents a future society, where personal genomics is readily available to anyone, and explores its societal impact.
See also
References
- ^ Toward Precision Medicine: Building a Knowledge Network for Biomedical Research and a New Taxonomy of Disease. The National Academies Press. 2011.
- ^ "The Pharmacogenomics Knowledgebase".
- ^ "NIH Office of Rare Disease Research".
- ^ "Gene Tests".
- ^ a b Wetterstrand, Kris (21 May 2012). "DNA Sequencing Costs: Data from the NHGRI Large-Scale Genome Sequencing Program". Large-Scale Genome Sequencing Program. National Human Genome Research Institute. Retrieved 24 May 2012.
- ^ "Coming Soon: Your Personal DNA Map?". News.nationalgeographic.com. 28 October 2010. Retrieved 19 October 2011.
- ^ "JDW-genome-supp-mat-march-proof.doc" (PDF). Retrieved 19 October 2011.
- ^ "True Single Molecule Sequencing (tSMS): Helicos BioSciences". Helicosbio.com. Retrieved 19 October 2011.
- ^ "Knome Lowers Price of Full Genome From $350,000 to $99,000". The Genetic Genealogist.
- ^ Karow, Julia (19 May 2009). "Knome Adds Exome Sequencing, Starts Offering Services to Researchers". GenomeWeb. Retrieved 24 February 2010.
- ^ Harmon, Katherine (28 June 2010). "Genome Sequencing for the Rest of Us". Scientific American. Retrieved 13 August 2010.
- ^ "Individual genome sequencing – Illumina, Inc". Everygenome.com. Retrieved 19 October 2011.
- ^ "Illumina Cutting Personal Genome Sequencing Price by 60% | GPlus.com". Glgroup.com. 4 June 2010. Retrieved 19 October 2011.
- ^ Karow, Julia. "Complete Genomics to Offer $5,000 Human Genome as a Service Business in Q2 2009 | In Sequence | Sequencing". GenomeWeb. Retrieved 19 October 2011.
- ^ Lauerman, John (5 February 2009). "Complete Genomics Drives Down Cost of Genome Sequence to $5,000". Bloomberg. Retrieved 19 October 2011.
- ^ [1][dead link]
- ^ "Illumina launches personal genome sequencing service for $48,000 : Genetic Future". Scienceblogs.com. Retrieved 19 October 2011.
- ^ {http://www.healthcarejournallr.com/the-journal/contents-index/features/563-what-a-tangled-web-we-weave.html}
- ^ https://sequencing.com/
- ^ https://sequencing.com/researcher/
- ^ https://sequencing.com/about-sequencingcom
- ^ https://sequencing.com/developer/
- ^ https://sequencing.com/altruist-endeavor
- ^ January 2009+BW20090112 "Illumina and Oxford Nanopore Enter into Broad Commercialization Agreement". Reuters. 12 January 2009. Retrieved 23 February 2009.
{{cite news}}: Check|url=value (help) - ^ http://www.genomeweb.com/sequenom-licenses-nanopore-technology-harvard-develop-third-generation-sequencer
- ^ "Single Molecule Real Time (SMRT) DNA Sequencing". Pacific Biosciences. Retrieved 23 February 2009. [dead link]
- ^ "Complete Human Genome Sequencing Technology Overview" (PDF). Complete Genomics. 2009. Retrieved 23 February 2009. [dead link]
- ^ http://files.shareholder.com/downloads/CRGN/0x0x53381/386c4aaa-f36e-4b7a-9ff0-c06e61fad31f/211559.pdf
- ^ "Genetics and Health Insurance State Anti-Discrimination Laws".
- ^ McCabe LL; McCabe ER (June 2001). "Postgenomic medicine. Presymptomatic testing for prediction and prevention". Clin Perinatol. 28 (2): 425–34. doi:10.1016/S0095-5108(05)70094-4. PMID 11499063.
- ^ Nelson RM; Botkjin JR; Kodish ED; et al. (June 2001). "Ethical issues with genetic testing in pediatrics". Pediatrics. 107 (6): 1451–5. doi:10.1542/peds.107.6.1451. PMID 11389275.
- ^ Borry P; Fryns JP; Schotsmans P; Dierickx K (February 2006). "Carrier testing in minors: a systematic review of guidelines and position papers". Eur. J. Hum. Genet. 14 (2): 133–8. doi:10.1038/sj.ejhg.5201509. PMID 16267502.
- ^ Borry P; Stultiens L; Nys H; Cassiman JJ; et al. (November 2006). "Presymptomatic and predictive genetic testing in minors: a systematic review of guidelines and position papers". Clin. Genet. 70 (5): 374–81. doi:10.1111/j.1399-0004.2006.00692.x. PMID 17026616.
- ^ Mark Johnson; Kathleen Gallagher (27 February 2011). "One in a Billion. Nic Volker case may be the leading edge of a wave moving across genetic medicine". Milwaukee Journal Sentinel.
{{cite news}}: Unknown parameter|last-author-amp=ignored (|name-list-style=suggested) (help) - ^ Lunshof, Jeantine; Mardis, Elaine. [Retrieved from http://www.future-science-group.com/_img/pics/Mardis_-_Foreward.pdf "Navigenics – How it works"] (PDF). Future Medicine Magazine. Retrieved 30 March 2012.
{{cite web}}: Check|url=value (help) - ^ "Genome: Unlocking Life's Code, Smithsonian Exhibit". Retrieved 7 June 2013.
- ^ Machácková, Eva (2003). "Disease-causing mutations versus neutral polymorphism: Use of bioinformatics and DNA diagnosis". Cas Lek Cesk. 142 (3). Czech Republic: Ceskoslovenska Lekarska Spolecnost: 150–153. PMID 12756842.
- ^ Hunter DJ; Khoury MJ; Drazen JM (January 2008). "Letting the genome out of the bottle—will we get our wish?". N. Engl. J. Med. 358 (2): 105–7. doi:10.1056/NEJMp0708162. PMID 18184955.
- ^ Lea DH; Skirton H; Read CY; Williams JK (March 2011). "Implications for educating the next generation of nurses on genetics and genomics in the 21st century". J Nurs Scholarsh. 43 (1): 3–12. doi:10.1111/j.1547-5069.2010.01373.x. PMID 21342419.
- ^ Gurwitz D; Bregman-Eschet Y (July 2009). "Personal genomics services: whose genomes?". Eur. J. Hum. Genet. 17 (7): 883–9. doi:10.1038/ejhg.2008.254. PMC 2986500. PMID 19259127.
- ^ De Cristofaro, E. (2012). "Whole Genome Sequencing: Innovation Dream or Privacy Nightmare?". ArXiv Repository.
- ^ Presidential Commission for the Study of Bioethical Issues (2012). "Privacy and Progress in Whole Genome Sequencing" (PDF).
{{cite journal}}: Cite journal requires|journal=(help) - ^ Blow N (October 2007). "Genomics: the personal side of genomics". Nature. 449 (7162): 627–30. doi:10.1038/449627a. PMID 17914399.
Bibliography
- Dudley; Karczewski (2013). Exploring Personal Genomics. Oxford University Press. ISBN 978-0199644490.
{{cite book}}: Unknown parameter|last-author-amp=ignored (|name-list-style=suggested) (help) - Sweet K; Michaelis R (May 2011). The Busy Physician's Guide to Genetics, Genomics and Personalized Medicine (1st ed.). Springer Scientific Press. ISBN 978-94-007-1147-1.
- Cadwalladr, Carole (8 June 2013). "What happened when I had my genome sequenced". The Guardian. Retrieved 10 July 2013.
