TRAIL
In the field of cell biology, TNF-related apoptosis-inducing ligand (TRAIL), is a protein functioning as a ligand that induces the process of cell death called apoptosis.[5][6]
TRAIL is a cytokine that is produced and secreted by most normal tissue cells. It causes apoptosis primarily in tumor cells,[7] by binding to certain death receptors. TRAIL and its receptors have been used as the targets of several anti-cancer therapeutics since the mid-1990s, such as Mapatumumab. However, as of 2013, these have not shown significant survival benefit.[8] TRAIL has also been implicated as a pathogenic or protective factor in various pulmonary diseases, particularly pulmonary arterial hypertension. [9]
TRAIL has also been designated CD253 (cluster of differentiation 253) and TNFSF10 (tumor necrosis factor (ligand) superfamily, member 10).[7]
Gene
In humans, the gene that encodes TRAIL is located at chromosome 3q26, which is not close to other TNF family members.[5] The genomic structure of the TRAIL gene spans approximately 20 kb and is composed of five exonic segments 222, 138, 42, 106, and 1245 nucleotides and four introns of approximately 8.2, 3.2, 2.3 and 2.3 kb.
The TRAIL gene lacks TATA and CAAT boxes and the promotor region contains putative response elements for transcription factors GATA, AP-1, C/EBP, SP-1, OCT-1, AP3, PEA3, CF-1, and ISRE.[citation needed]
The TRAIL gene as a drug target
TIC10 (which causes expression of TRAIL) was investigated in mice with various tumour types.[8]
Small molecule ONC201 causes expression of TRAIL which kills some cancer cells.[10]
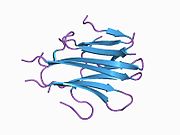
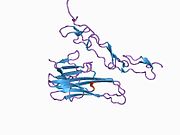
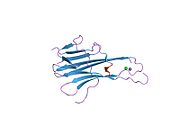
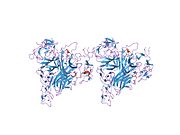
Structure
TRAIL shows homology to other members of the tumor necrosis factor superfamily. It is composed of 281 amino acids and has characteristics of a type II transmembrane protein (i.e. no leader sequence and an internal transmembrane domain). The N-terminal cytoplasmic domain is not conserved across family members, however, the C-terminal extracellular domain is conserved and can be proteolytically cleaved from the cell surface. TRAIL forms a homotrimer that binds three receptor molecules.
Function
TRAIL binds to the death receptors DR4 (TRAIL-RI) and DR5 (TRAIL-RII). The process of apoptosis is caspase-8-dependent. Caspase-8 activates downstream effector caspases including procaspase-3, -6, and -7, leading to activation of specific kinases.[11] TRAIL also binds the receptors DcR1 and DcR2, which do not contain a cytoplasmic domain (DcR1) or contain a truncated death domain (DcR2). DcR1 functions as a TRAIL-neutralizing decoy-receptor. The cytoplasmic domain of DcR2 is functional and activates NFkappaB. In cells expressing DcR2, TRAIL binding therefore activates NFkappaB, leading to transcription of genes known to antagonize the death signaling pathway and/or to promote inflammation. Application of engineered ligands that have variable affinity for different death (DR4 and DR5) and decoy receptors (DCR1 and DCR2) may allow selective targeting of cancer cells by controlling activation of type1/type 2 pathways of cell death and single cell fluctuations.
The TRAIL receptors as a drug target
In clinical trials only a small proportion of patients responded to various drugs that targeted TRAIL death receptors.[12]
Interactions
TRAIL has been shown to interact with TNFRSF10B.[13][14][15]
See also
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000121858 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000039304 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b Wiley SR, Schooley K, Smolak PJ, Din WS, Huang CP, Nicholl JK, Sutherland GR, Smith TD, Rauch C, Smith CA (December 1995). "Identification and characterization of a new member of the TNF family that induces apoptosis". Immunity. 3 (6): 673–82. doi:10.1016/1074-7613(95)90057-8. PMID 8777713.
- ^ Pitti RM, Marsters SA, Ruppert S, Donahue CJ, Moore A, Ashkenazi A (May 1996). "Induction of apoptosis by Apo-2 ligand, a new member of the tumor necrosis factor cytokine family". J. Biol. Chem. 271 (22): 12687–90. doi:10.1074/jbc.271.22.12687. PMID 8663110.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ a b "TNFSF10". NCBI Gene.
- ^ a b Cormier Z (February 2013). "Small-molecule drug drives cancer cells to suicide". Nature. 494. doi:10.1038/nature.2013.12385.
- ^ Braithwaite, Adam T.; Marriott, Helen M.; Lawrie, Allan (2018). "Divergent Roles for TRAIL in Lung Diseases". Frontiers in Medicine. 5. doi:10.3389/fmed.2018.00212. ISSN 2296-858X.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ ONC201: Stressing tumors to death. Feb 2016
- ^ Song JJ, Lee YJ (May 2008). "Differential cleavage of Mst1 by caspase-7/-3 is responsible for TRAIL-induced activation of the MAPK superfamily". Cell. Signal. 20 (5): 892–906. doi:10.1016/j.cellsig.2008.01.001. PMC 2483832. PMID 18276109.
- ^ Dimberg, L Y; Anderson, C K; Camidge, R; Behbakht, K; Thorburn, A; Ford, H L (14 March 2013). "On the TRAIL to successful cancer therapy? Predicting and counteracting resistance against TRAIL-based therapeutics". Oncogene. 32 (11): 1341–1350. doi:10.1038/onc.2012.164. PMC 4502956.
- ^ Kaptein A, Jansen M, Dilaver G, Kitson J, Dash L, Wang E, Owen MJ, Bodmer JL, Tschopp J, Farrow SN (November 2000). "Studies on the interaction between TWEAK and the death receptor WSL-1/TRAMP (DR3)". FEBS Lett. 485 (2–3): 135–41. doi:10.1016/S0014-5793(00)02219-5. PMID 11094155.
- ^ Walczak H, Degli-Esposti MA, Johnson RS, Smolak PJ, Waugh JY, Boiani N, Timour MS, Gerhart MJ, Schooley KA, Smith CA, Goodwin RG, Rauch CT (September 1997). "TRAIL-R2: a novel apoptosis-mediating receptor for TRAIL". EMBO J. 16 (17): 5386–97. doi:10.1093/emboj/16.17.5386. PMC 1170170. PMID 9311998.
- ^ Hymowitz SG, Christinger HW, Fuh G, Ultsch M, O'Connell M, Kelley RF, Ashkenazi A, de Vos AM (October 1999). "Triggering cell death: the crystal structure of Apo2L/TRAIL in a complex with death receptor 5". Mol. Cell. 4 (4): 563–71. doi:10.1016/S1097-2765(00)80207-5. PMID 10549288.
Further reading
- Almasan A, Ashkenazi A (2004). "Apo2L/TRAIL: apoptosis signaling, biology, and potential for cancer therapy". Cytokine Growth Factor Rev. 14 (3–4): 337–48. doi:10.1016/S1359-6101(03)00029-7. PMID 12787570.
- Cha SS, Song YL, Oh BH (2004). "Specificity of molecular recognition learned from the crystal structures of TRAIL and the TRAIL:sDR5 complex". Vitam. Horm. Vitamins & Hormones. 67: 1–17. doi:10.1016/S0083-6729(04)67001-4. ISBN 978-0-12-709867-8. PMID 15110168.
- Song C, Jin B (2005). "TRAIL (CD253), a new member of the TNF superfamily". J. Biol. Regul. Homeost. Agents. 19 (1–2): 73–7. PMID 16178278.
- Bucur O, Ray S, Bucur MC, Almasan A (2006). "APO2 ligand/tumor necrosis factor-related apoptosis-inducing ligand in prostate cancer therapy". Front. Biosci. 11: 1549–68. doi:10.2741/1903. PMID 16368536.
External links
- [1] Apoptosis, Trail & Caspase 8 - The Proteolysis Map-animation
- PDB: 1D2Q
- TRAIL+Protein at the U.S. National Library of Medicine Medical Subject Headings (MeSH)